// CERCETARE
OncoGen
Transformăm știința în speranță! De la terapii genice celulare și imunoterapii anti-cancer, până la bioimprimarea 3D a țesuturilor tumorale, descoperirile echipei OncoGen sunt publicate în reviste internaționale de top și recunoscute în consorții europene finanțate de UE. Medicina viitorului se construiește în Timișoara!
Galerie Video
Explorați prezentări, conferințe și documentare despre cercetarea noastră
Prezentarea OncoGen
Descoperă centrul de cercetare și facilități
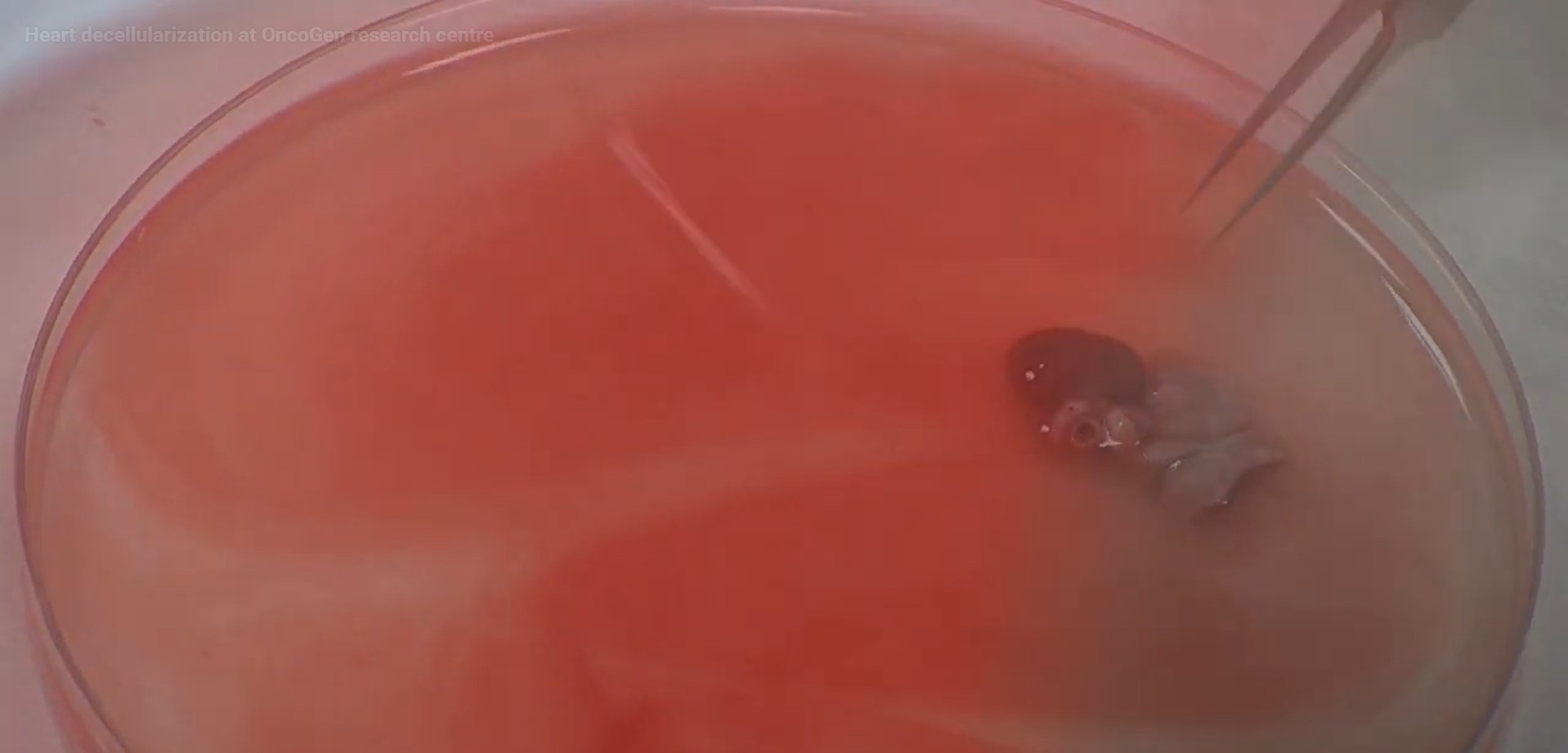
Heart decellularization at OncoGen research centre
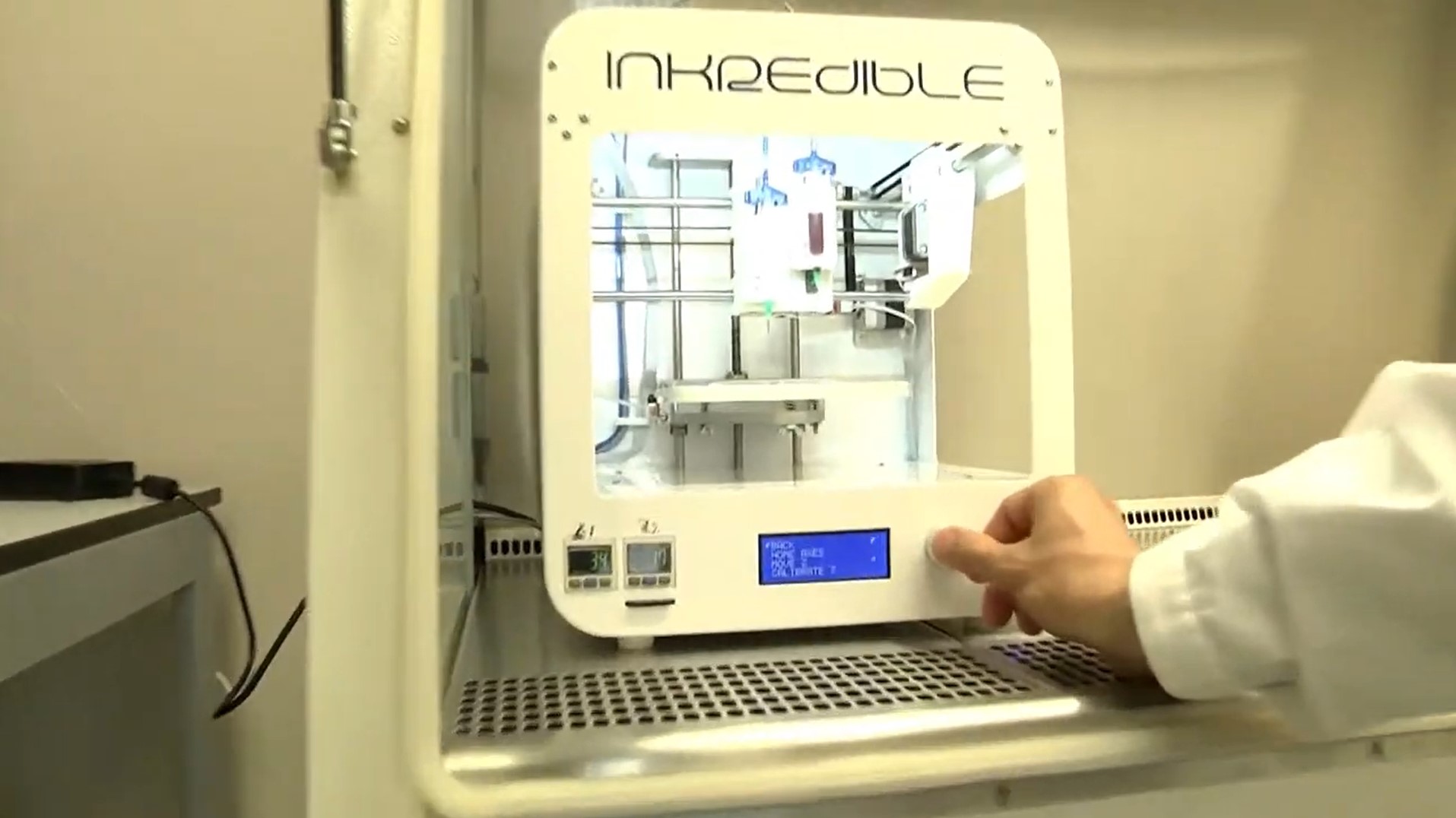

3D printing blood vessel at OncoGen research centre
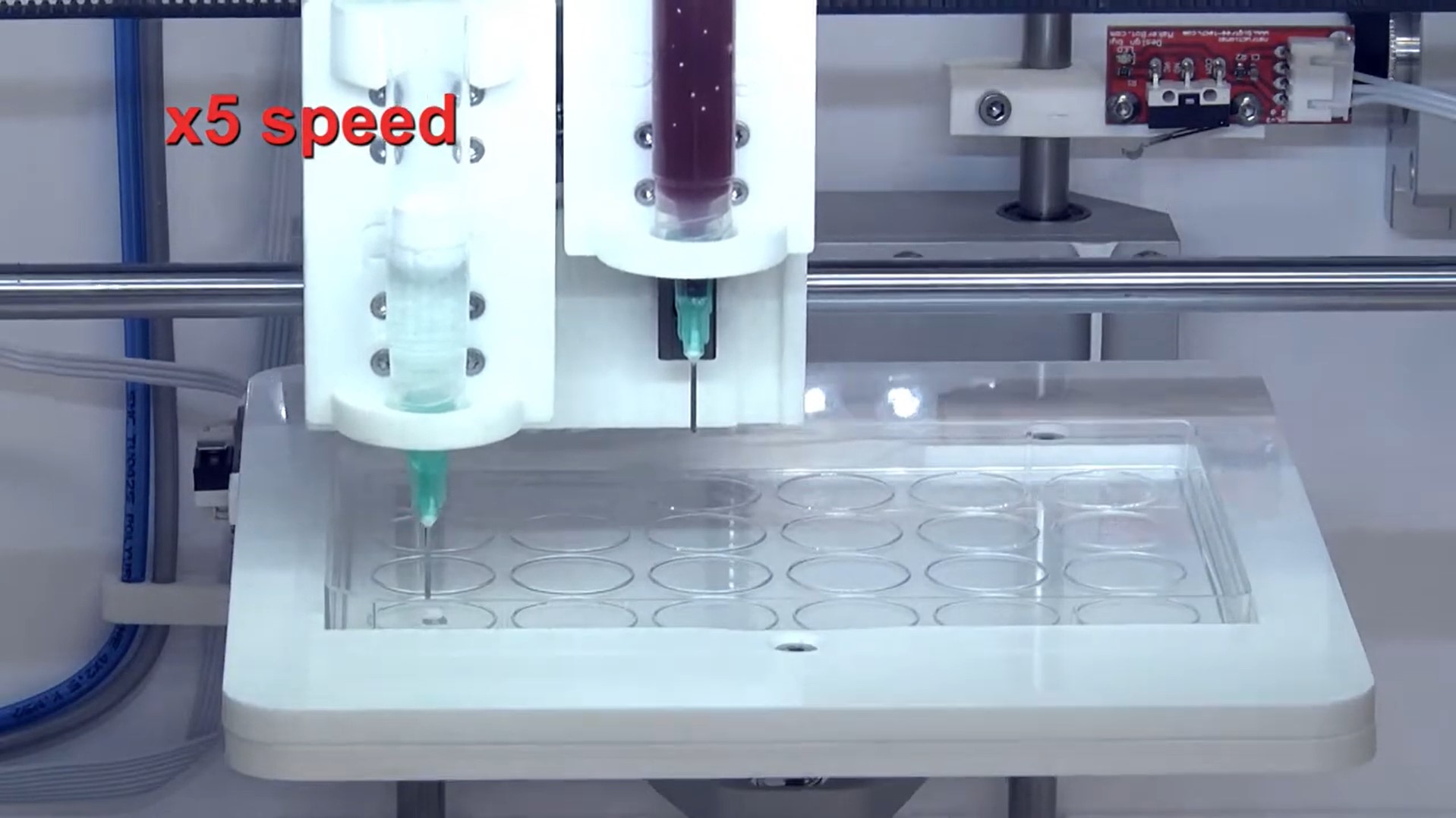

3D printing tumors at OncoGen research centre

S-a terminat pandemia? Subestimăm virusul?
Concluziile după doi ani | prof. Virgil Păunescu, OncoGen
Principalele proiecte de cercetare dezvoltate la OncoGen
Explorați selecta noastră de proiecte finanțate de Uniunea Europeană și instituții internaționale
DECODE
Vezi detalii →INSPIRED
Vezi detalii →ROSECAN
Vezi detalii →DeLIMIT
Vezi detalii →NANOCEL
Vezi detalii →Știri și Evenimente
COMUNICAT DE PRESĂ: Centrul OncoGen participă în proiectul TheraFast
Centrul OncoGen anunță participarea sa în proiectul TheraFast, o inițiativă inovatoare pentru terapiile de genie genetic.
Citește mai mult →Centrul de Terapii Genice și Celulare OncoGen anunță începerea proiectului DECODEX
Proiect revoluționar în domeniul terapiilor genice și celulare, marcat de inovații în diagnosticul și tratamentul personalizat.
Citește mai mult →Digipedia: AI poate schimba vieți
OncoGen participă la conferința Digipedia, explorând rolul inteligenței artificiale în transformarea domeniului medical și cercetării.
Citește mai mult →